Related News
Copyright 2012 neutronsources.org | All rights reserved. | Powered by FRM II | Imprint / Privacy Policy
Catching a glimpse at enzymes on the job: a combined time-resolved neutron scattering and fluorescence study
Date: 15/02/2017
Source: ill.eu
AAA+ ATPases are a large family of ubiquitous enzymes with multiple tasks, including the remodelling of the cellular proteome, i.e. the ensemble of proteins in a biological cell. A subfamily, so-called unfoldases, recognize, unfold, and address misfolded or dysfunctional proteins towards proteolytic complexes which eliminate these potentially toxic proteins in order to assure a healthy, functional state of the cellular proteome. Given the intrinsic flexibility of ATPases and the transient character of the interaction with their protein substrates, it is challenging in structural biology to follow the conformational changes of these enzyme-substrate complexes during the active unfolding process.
In a collaboration between ILL and IBS a novel approach was developed combining time-resolved small angle neutron scattering (TR-SANS) with online-fluorescence spectroscopy on D22 in order to monitor the PAN unfoldase from the deep-sea Methanocaldococcus jannaschii organism and a Green Fluorescent Protein (GFP) model substrate. By using alternating perdeuteration of both partners and by controlling the enzymatic activity of the hyperthermophilic PAN system by temperature activation at 55-60 °C, it was possible to follow conformational changes of both PAN and GFP separately and individually during the active unfolding process at a time resolution of 30 seconds.
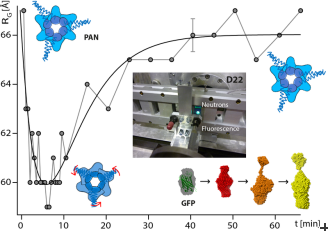
The results show the progressive unfolding and aggregation of GFP as well as a concomitant and reversible contraction of the PAN unfoldase during the active reaction (Fig. 1). While the methodological approach developed has been designed for this specific project, it is expected that it can be applied to a wide range of biological macromolecular complexes and provide structural information from individual partners at a time resolution of some seconds.
Original Publication
Re.: Scientific Reports 7, Article number: 40948 (2017) doi:10.1038/srep40948
Contact: Dr F. Gabel, IBS & ILL